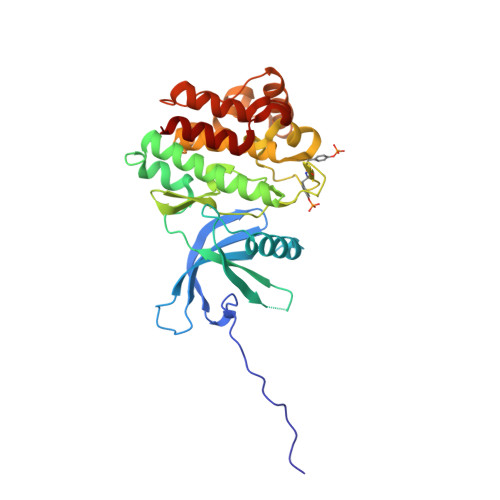
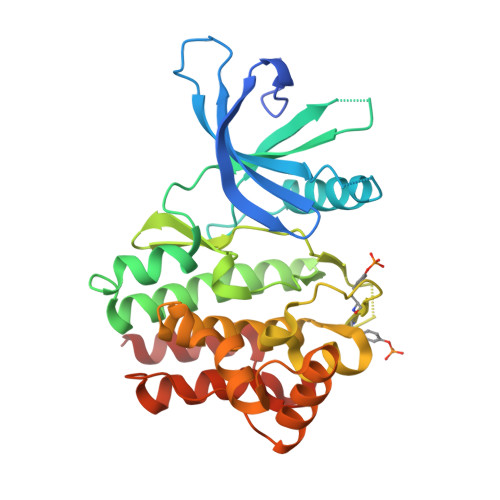
Discovery of (2R)-N-[3-[2-[(3-Methoxy-1-methyl-pyrazol-4-yl)amino]pyrimidin-4-yl]-1H-indol-7-yl]-2-(4-methylpiperazin-1-yl)propenamide (AZD4205) as a Potent and Selective Janus Kinase 1 Inhibitor.
Su, Q., Banks, E., Bebernitz, G., Bell, K., Borenstein, C.F., Chen, H., Chuaqui, C.E., Deng, N., Ferguson, A.D., Kawatkar, S., Grimster, N.P., Ruston, L., Lyne, P.D., Read, J.A., Peng, X., Pei, X., Fawell, S., Tang, Z., Throner, S., Vasbinder, M.M., Wang, H., Winter-Holt, J., Woessner, R., Wu, A., Yang, W., Zinda, M., Kettle, J.G.(2020) J Med Chem 63: 4517-4527
- PubMed: 32297743
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01392
- Primary Citation of Related Structures:
6SM8, 6SMB - PubMed Abstract:
JAK1, JAK2, JAK3, and TYK2 belong to the JAK (Janus kinase) family. They play critical roles in cytokine signaling. Constitutive activation of JAK/STAT pathways is associated with a wide variety of diseases. Particularly, pSTAT3 is observed in response to the treatment with inhibitors of oncogenic signaling pathways such as EGFR, MAPK, and AKT and is associated with resistance or poorer response to agents targeting these pathways. Among the JAK family kinases, JAK1 has been shown to be the primary driver of STAT3 phosphorylation and signaling; therefore, selective JAK1 inhibition can be a viable means to overcome such treatment resistances. Herein, an account of the medicinal chemistry optimization from the promiscuous kinase screening hit 3 to the candidate drug 21 (AZD4205), a highly selective JAK1 kinase inhibitor, is reported. Compound 21 has good preclinical pharmacokinetics. Compound 21 displayed an enhanced antitumor activity in combination with an approved EGFR inhibitor, osimertinib, in a preclinical non-small-cell lung cancer (NSCLC) xenograft NCI-H1975 model.
Organizational Affiliation:
Oncology R&D, AstraZeneca R&D, Waltham, Massachusetts 02451, United States.